Geopandas¶
Welcome to the next level: Level 5 - spyndex + geopandas!
Remember to install spyndex!
And also remember to install
geopandas! I recommend you to install it using conda!
[ ]:
!pip install -U spyndex
Now, let’s start!
First, import spyndex and geopandas:
[56]:
import spyndex
import geopandas as gpd
geopandas.GeoDataFrame¶
In geopandas, each column is a pandas.Series. The exception is the geometry column, which is a geopandas.GeoSeries.
Here we are going to compute spectral indices using pandas.Series data types, but adding them to a geopandas.GeoDataFrame, making them possible to map!
First, let’s create a random sample of reflectances from Sentinel-2. For this, we are going to use Google Earth Engine. Let’s install some useful packages:
[ ]:
!pip install eemont geemap
Then, let’s import them and initialize Earth Engine.
[57]:
import ee, eemont, geemap
Map = geemap.Map()
Now, let’s use the first image in the Sentinel-2 SR collection and sample 1000 random points from it.
With the
eemontpackage we can mask clouds and scale and offset the image!
[58]:
poi = ee.ImageCollection("COPERNICUS/S2_SR") \
.first() \
.maskClouds() \
.scaleAndOffset() \
.sample(numPixels = 1000,geometries = True)
Now, let’s use geemap to convert the samples to a geopandas.GeoDataFrame:
[59]:
gdf = geemap.ee_to_geopandas(poi)
You can see that the geopandas.GeoDataFrame has the geometry column and the remaining columns are the samples from all bands in the image!
[60]:
gdf.head()
[60]:
| geometry | AOT | B1 | B11 | B12 | B2 | B3 | B4 | B5 | B6 | ... | B8A | B9 | QA10 | QA20 | QA60 | SCL | TCI_B | TCI_G | TCI_R | WVP | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 0 | POINT (27.29076 26.40036) | 0.197 | 0.1070 | 0.6306 | 0.5504 | 0.1367 | 0.2429 | 0.3712 | 0.4120 | 0.4322 | ... | 0.4331 | 0.4570 | 0 | 0 | 0 | 5 | 139 | 248 | 255 | 0.695 |
| 1 | POINT (28.06818 26.58967) | 0.198 | 0.1697 | 0.5636 | 0.4815 | 0.2111 | 0.3039 | 0.3790 | 0.4122 | 0.4194 | ... | 0.4274 | 0.4324 | 0 | 0 | 0 | 5 | 215 | 255 | 255 | 0.815 |
| 2 | POINT (27.87679 26.33568) | 0.193 | 0.1670 | 0.6501 | 0.5622 | 0.2118 | 0.3114 | 0.4240 | 0.4666 | 0.4760 | ... | 0.4990 | 0.5049 | 0 | 0 | 0 | 5 | 216 | 255 | 255 | 0.644 |
| 3 | POINT (27.04346 26.21331) | 0.195 | 0.1363 | 0.7335 | 0.6691 | 0.1859 | 0.3268 | 0.4842 | 0.5332 | 0.5514 | ... | 0.5692 | 0.5823 | 0 | 0 | 0 | 5 | 189 | 255 | 255 | 0.624 |
| 4 | POINT (27.53605 26.84894) | 0.194 | 0.2098 | 0.6491 | 0.5494 | 0.2606 | 0.3834 | 0.5202 | 0.5680 | 0.5758 | ... | 0.5684 | 0.5798 | 0 | 0 | 0 | 5 | 255 | 255 | 255 | 0.797 |
5 rows × 22 columns
Now, let’s check the indices to compute: The IRECI and the NDVI:
[61]:
spyndex.indices.IRECI
[61]:
IRECI: Inverted Red-Edge Chlorophyll Index (attributes = ['bands', 'contributor', 'date_of_addition', 'formula', 'long_name', 'reference', 'short_name', 'type'])
[62]:
spyndex.indices.NDVI
[62]:
NDVI: Normalized Difference Vegetation Index (attributes = ['bands', 'contributor', 'date_of_addition', 'formula', 'long_name', 'reference', 'short_name', 'type'])
[63]:
spyndex.indices.IRECI.bands
[63]:
('RE3', 'R', 'RE1', 'RE2')
[64]:
spyndex.indices.NDVI.bands
[64]:
('N', 'R')
We need the Red Edge bands from Sentinel-2 plus the Red and NIR bands!
Let’s make a list of the indices:
[65]:
indicesToCompute = ["IRECI","NDVI"]
Now, let’s compute the indices and add them directly to our geopandas.GeoDataFrame:
[66]:
gdf[indicesToCompute] = spyndex.computeIndex(
index = indicesToCompute,
params = {
"R": gdf["B4"],
"RE1": gdf["B5"],
"RE2": gdf["B6"],
"RE3": gdf["B7"],
"N": gdf["B8"]
}
)
You can see that the indices were added as new columns!
[67]:
gdf.head()
[67]:
| geometry | AOT | B1 | B11 | B12 | B2 | B3 | B4 | B5 | B6 | ... | QA10 | QA20 | QA60 | SCL | TCI_B | TCI_G | TCI_R | WVP | IRECI | NDVI | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 0 | POINT (27.29076 26.40036) | 0.197 | 0.1070 | 0.6306 | 0.5504 | 0.1367 | 0.2429 | 0.3712 | 0.4120 | 0.4322 | ... | 0 | 0 | 0 | 5 | 139 | 248 | 255 | 0.695 | 0.073747 | 0.097496 |
| 1 | POINT (28.06818 26.58967) | 0.198 | 0.1697 | 0.5636 | 0.4815 | 0.2111 | 0.3039 | 0.3790 | 0.4122 | 0.4194 | ... | 0 | 0 | 0 | 5 | 215 | 255 | 255 | 0.815 | 0.045786 | 0.070281 |
| 2 | POINT (27.87679 26.33568) | 0.193 | 0.1670 | 0.6501 | 0.5622 | 0.2118 | 0.3114 | 0.4240 | 0.4666 | 0.4760 | ... | 0 | 0 | 0 | 5 | 216 | 255 | 255 | 0.644 | 0.066105 | 0.091688 |
| 3 | POINT (27.04346 26.21331) | 0.195 | 0.1363 | 0.7335 | 0.6691 | 0.1859 | 0.3268 | 0.4842 | 0.5332 | 0.5514 | ... | 0 | 0 | 0 | 5 | 189 | 255 | 255 | 0.624 | 0.085109 | 0.097904 |
| 4 | POINT (27.53605 26.84894) | 0.194 | 0.2098 | 0.6491 | 0.5494 | 0.2606 | 0.3834 | 0.5202 | 0.5680 | 0.5758 | ... | 0 | 0 | 0 | 5 | 255 | 255 | 255 | 0.797 | 0.061128 | 0.060926 |
5 rows × 24 columns
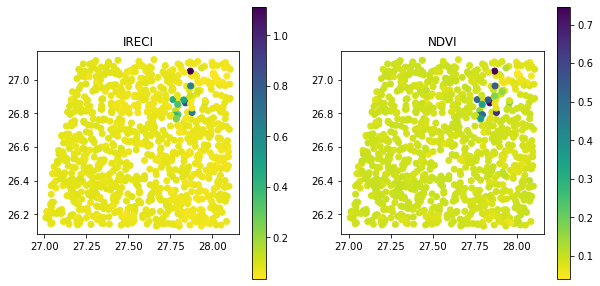
Finally, let’s map the indices!
[68]:
import matplotlib.pyplot as plt
You’ll see a lot of low values for both indices, that’s because that image is in the desert with some pivot crops!
[69]:
fig, ax = plt.subplots(1, 2,figsize = (10,5))
gdf.plot(column = 'IRECI',legend = True,ax = ax[0],cmap = "viridis_r")
ax[0].set(title = "IRECI")
gdf.plot(column = 'NDVI',legend = True,ax = ax[1],cmap = "viridis_r")
ax[1].set(title = "NDVI")
[69]:
[Text(0.5, 1.0, 'NDVI')]
That’s all for now! :) See you in a next tutorial!
